

The Future Is Here
Filter like a Pro with the most advanced eDNA sampling system on the market Buy Now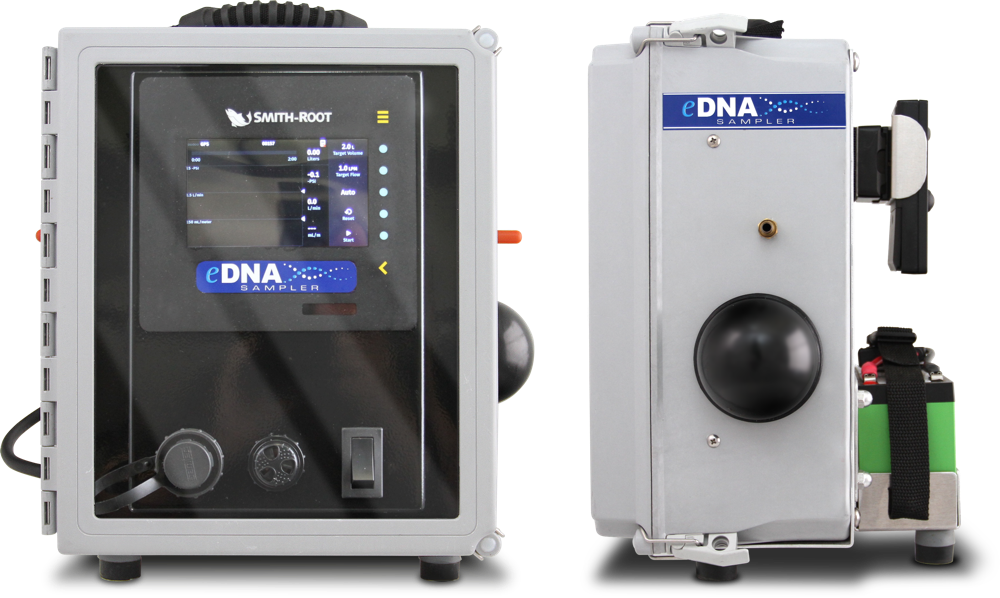
Standardize your workflow
Our computer-controlled pump makes sampling more replicable with user-defined sampling parameters
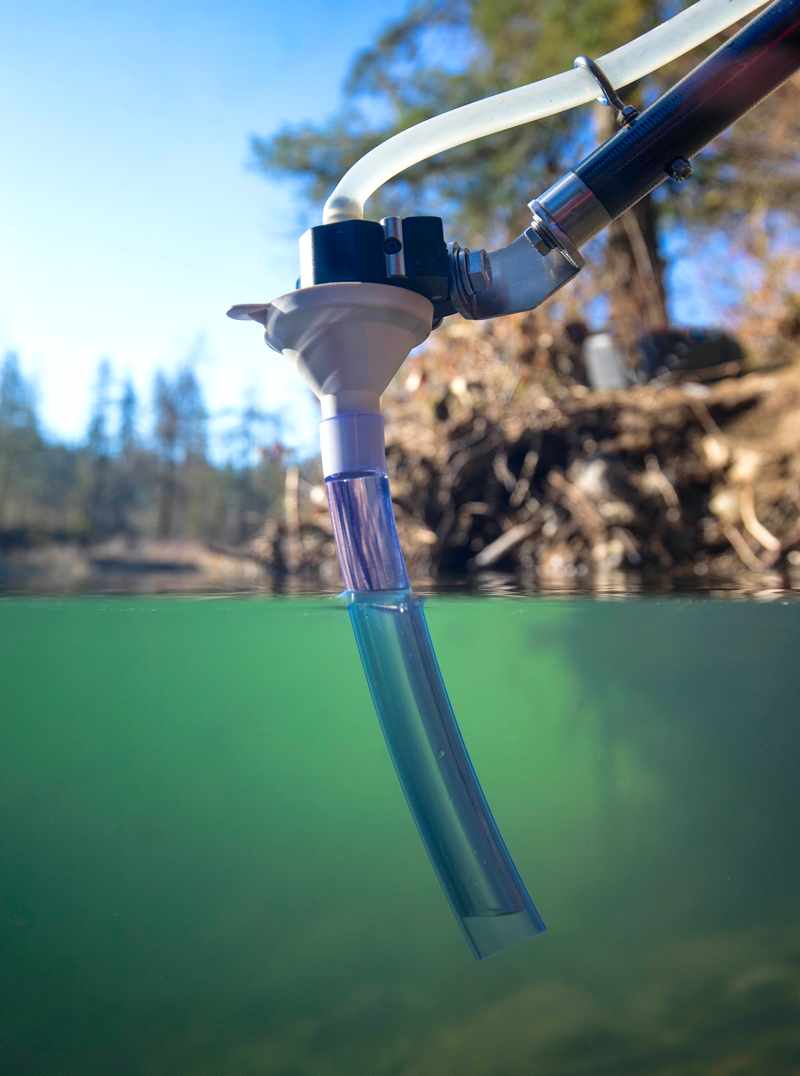

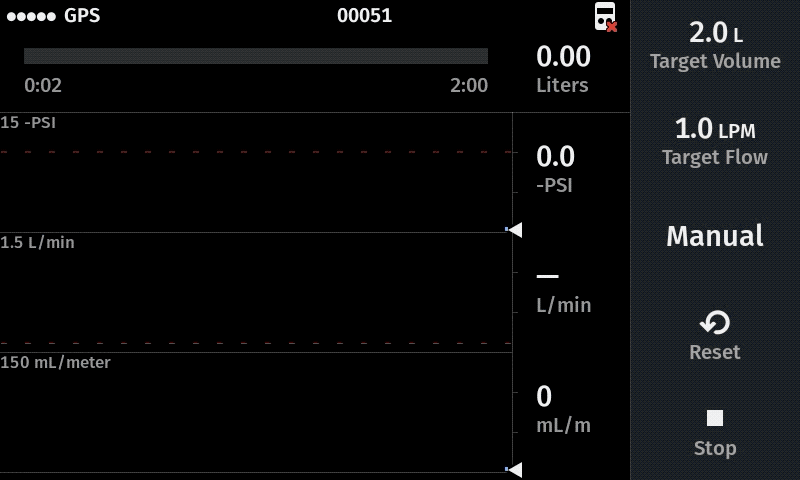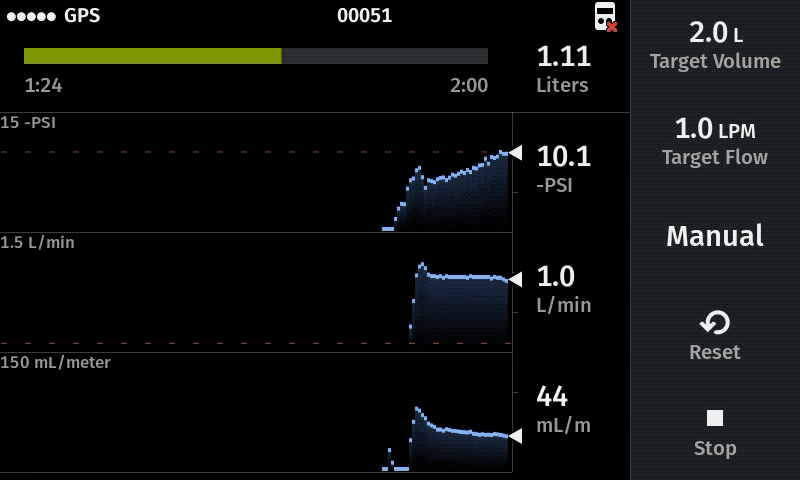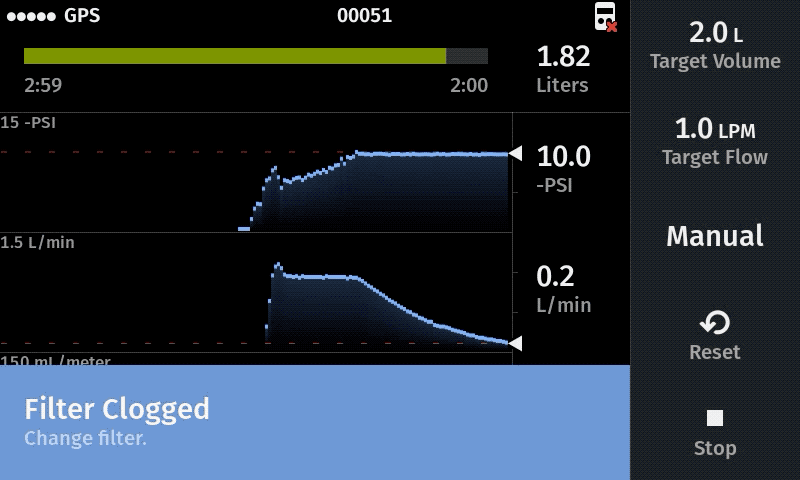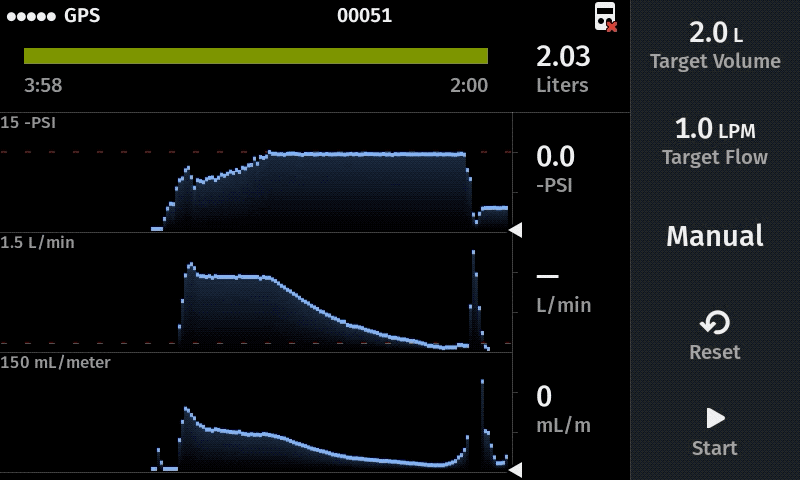

Improve sampling efficiency
Maximize species detection with Smith-Root eDNA filters
Filter rapidly for best use of field time
Minimize risk of contamination with a published method
Order FiltersOptimized for mobile sampling
eDNA is not standing still and neither should you. We designed the backpack model for continuous, mobile eDNA sampling. Generate spatially-integrated eDNA samples for maximal species detection.
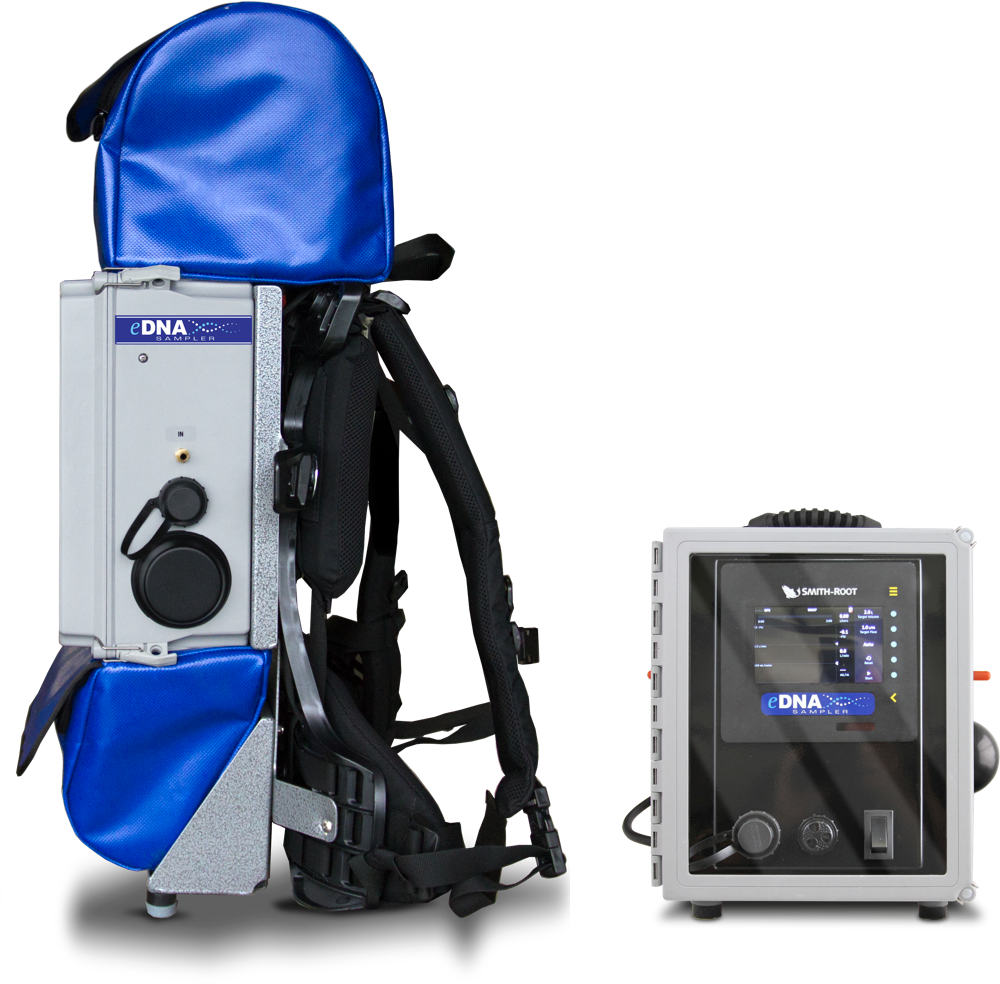

Customize for your application
Multiple filter pore sizes available
Regular and self-preserving eDNA filters
Telescoping pole and trident pole options
Lithium-iron and lead-acid batteries
Supported by science
Our team of scientists and engineers work continuously to provide the best possible support for the Smith-Root eDNA Sampler user community.
Software updates are issued regularly, and units can be upgraded remotely via the USB port.
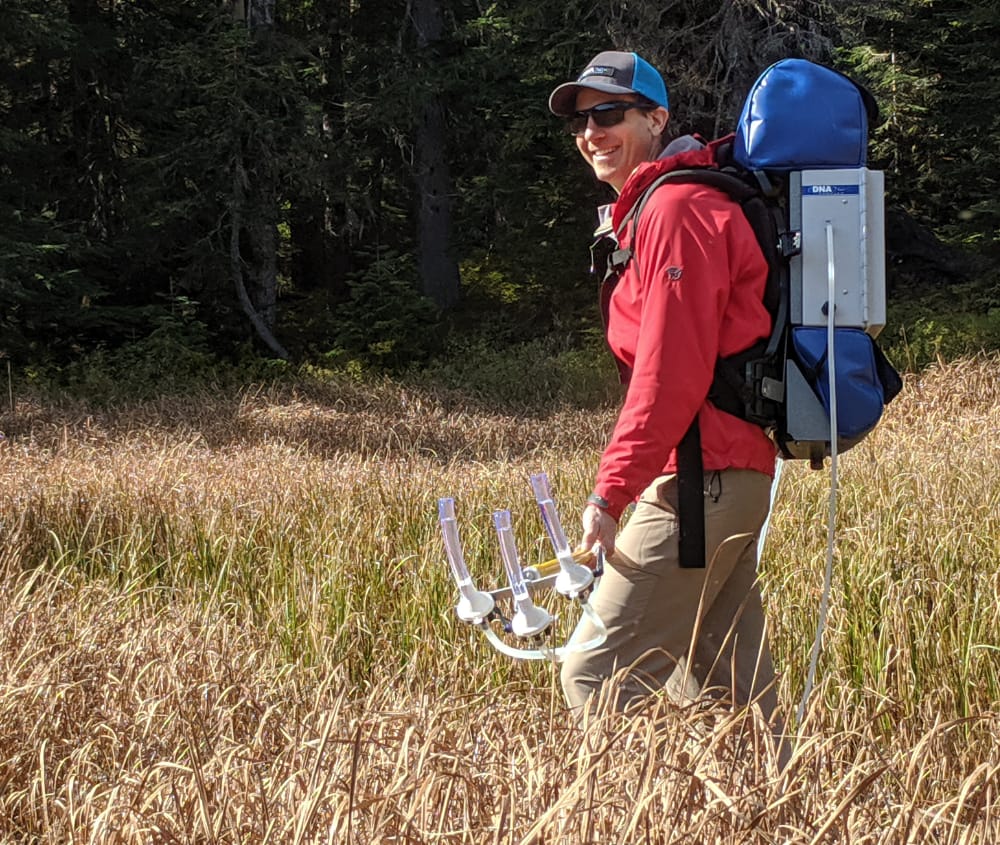

Research publications with eDNA Sampler
Gasparini, L., Crookes, S., Prosser, R.S. & Hanner, R. (2020) Detection of freshwater mussels (Unionidae) using environmental DNA in riverine systems. Environmental DNA, 2, 321-329.
Pope, K.L., Goldberg, C.S., Nelson, N.L., Cummings, A.K., Seaborn, T. & Piovia‐Scott, J. (2020) Designing environmental DNA surveys in complex aquatic systems: Backpack sampling for rare amphibians in Sierra Nevada meadows. Aquatic Conservation: Marine and Freshwater Ecosystems, 30, 1975-1987.
Skinner, M., Murdoch, M., Loeza-Quintana, T., Crookes, S. & Hanner, R. (2020) A mesocosm comparison of laboratory-based and on-site eDNA solutions for detection and quantification of striped bass (Morone saxatilis) in marine ecosystems. Environmental DNA, 2, 298-308.
Thomas, A.C., Howard, J., Nguyen, P.L., Seimon, T.A. & Goldberg, C.S. (2018) eDNA Sampler: A fully integrated environmental DNA sampling system. Methods in Ecology and Evolution, 9, 1379-1385.
Thomas, A.C., Nguyen, P.L., Howard, J. & Goldberg, C.S. (2019) A self‐preserving, partially biodegradable eDNA filter. Methods in Ecology and Evolution, 10, 1136-1141.
Thomas, A.C., Tank, S., Nguyen, P.L., Ponce, J., Sinnesael, M. & Goldberg, C.S. (2020) A system for rapid eDNA detection of aquatic invasive species. Environmental DNA, 2, 261-270.
FAQs
The pore size to use will depend on the organism you want to detect and the level of suspended particulate in the water you are sampling.
0.45 µm = Microbial studies such as fish pathogen detection or microbiome analysis.
1.2 – 5.0 µm = Fish, macroinvertebrates, bivalves, etc. (Metazoan studies).
We generally recommend 5 µm filters because they work well in a wide range of sampling habitats. 5.0 µm filters will actually capture slightly less eDNA per liter of water collected than 1.2 µm filters, but 5.0 µm usually allows for a much larger water volume to be filtered prior to clogging. The net effect is that 5.0µm filters will capture more total eDNA if the water volume is maximized.
Yes. We designed the Smith-Root eDNA filter housing to be compatible with a wide range of filter membrane materials. Please contact us if you do not see the pore size or material you want to use. Minimum orders may apply.
A wide range of sample water volumes have been used in the eDNA literature. Generally speaking, the goal is to maximize water volume per sample to increase species detection sensitivity, and also to standardize the water collected per filter sample within a study. However, the amount of water that can be collected at given pore size varies substantially between different habitats. A pilot study can be done to determine the realistic sample volume for your study system (please see the eDNA Sampler manual for details).
Filter samples of 1L – 5L are common water volumes for the eDNA Sampler system. We also recommend a minimum of 0.5 L per sample for the Smith-Root eDNA Sampler. If you cannot achieve 0.5 L per filter sample, we recommend increasing the eDNA Sampler pressure threshold or increasing the filter pore size.
New tools are currently in development to help with eDNA study design. The required number of samples will depend on the desired species detection probability, eDNA assay sensitivity, environmental factors, and other technical details. If you are interested in using the Smith-Root eDNA lab services, we can offer you guidance on study design in collaboration with our partner laboratory Precision Biomonitoring.
Online resources are also available for estimating your species detection probability under different sampling designs and assay variables. https://edna-probability-of-detection-calculator.shinyapps.io/eDNA_PrDetection_2/
For new studies we recommend a flow rate of 1.0 L/min as a starting point. When the water has a very high particulate load, a lower flow rate is recommended (e.g. 0.5 L/min) to help maximize the volume filtered prior to clogging. Higher flow rates (e.g 1.4 L/min) can be advisable when using multiple filters in the system (triplicate sampling) because the flow rate at each filter equals the displayed value divided by the total number of filters.
eDNA Sampler Specifications
| Sensors | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Minimum Flow Rate | 0.1 L/minute | ||||||||||||||||||
| Maximum Flow Rate | 1.4 L/minute | ||||||||||||||||||
| Maximum Measurable Volume | 999.9 L | ||||||||||||||||||
| Volume Accuracy | > 90% at 0.1 L/minute or higher flow | ||||||||||||||||||
| GPS System Compatibility | GNSS (GPS, Galileo, GLONASS, BeiDou) | ||||||||||||||||||
| Number Of Channels | 72 | ||||||||||||||||||
| GPS, GLONASS Sensitivity | -167 dBm | ||||||||||||||||||
| GPS Position Accuracy | 2.5 meter | ||||||||||||||||||
| Pump | |||||||||||||||||||
| Ports | 0.25 inch ID Tubing | ||||||||||||||||||
| Priming Vacuum | Approx. 5 psi | ||||||||||||||||||
| Maximum Wetted Vacuum | 14 psi (96.5 kPa) | ||||||||||||||||||
| Communications | |||||||||||||||||||
| USB | 2.0 | ||||||||||||||||||
| Log File Format | Comma Separated Values (csv) | ||||||||||||||||||
| Batteries | |||||||||||||||||||
| Sampler Battery | 11.1Ah 12.8V LiFePO4 | ||||||||||||||||||
| Sampler Battery Life | 6 to 8 hours | ||||||||||||||||||
| Remote Control Battery | 1900mAH 1.2V NiMH | ||||||||||||||||||
| Remote Control Battery Life | Approx. 4 hours | ||||||||||||||||||
| Size and Weight | |||||||||||||||||||
| Sampler Size and Weight | Height: 16 in. (40.64 cm) Width: 15 in. (38.1 cm) Depth: 9.25 in. (23.5 cm) Weight: 16.2 lb. (7.35 kg) without battery or remote control |
||||||||||||||||||
| Remote Control Size and Weight | Height: 4.7 in. (11.94 cm) Width: 2.5 in. (6.35 cm) Depth: 0.94 in. (2.39 cm) Weight: 0.4 lb. (181.4 g) |
||||||||||||||||||
| Remote Control Battery Weight | 0.1 lb. (45.4 g) | ||||||||||||||||||
Specifications subject to change without notice.